With an average weight of ~12 kg and surface area of ~2m², skin is the largest organ of the body and is made up of three layers: epidermis, dermis, and hypodermis. Epidermis, the outermost layer, is composed of densely packed epithelial cells. Under the epidermis lies the dermis, which mainly contains blood vessels, hair follicles and sweat glands. The third layer, the hypodermis, is composed of loose connective and fatty tissue (Fig. 1). Given its large area and exposure to external elements, the skin is susceptible to injury, ranging from minor bruises to cuts, lacerations, and tears. The skin’s response to insults like these is a process well-known as wound healing, which occurs in three stages: inflammation, proliferation, and tissue remodeling. These stages work together in an overlapping sequential manner to ensure complete wound healing. A failure in any of the normal wound healing stages leads to chronic wounding and aberrant scar formation as seen in burn injuries and scar tissue formations. A delay in wound healing can result in increased infections and permanent tissue damage as seen in patients with diabetes.
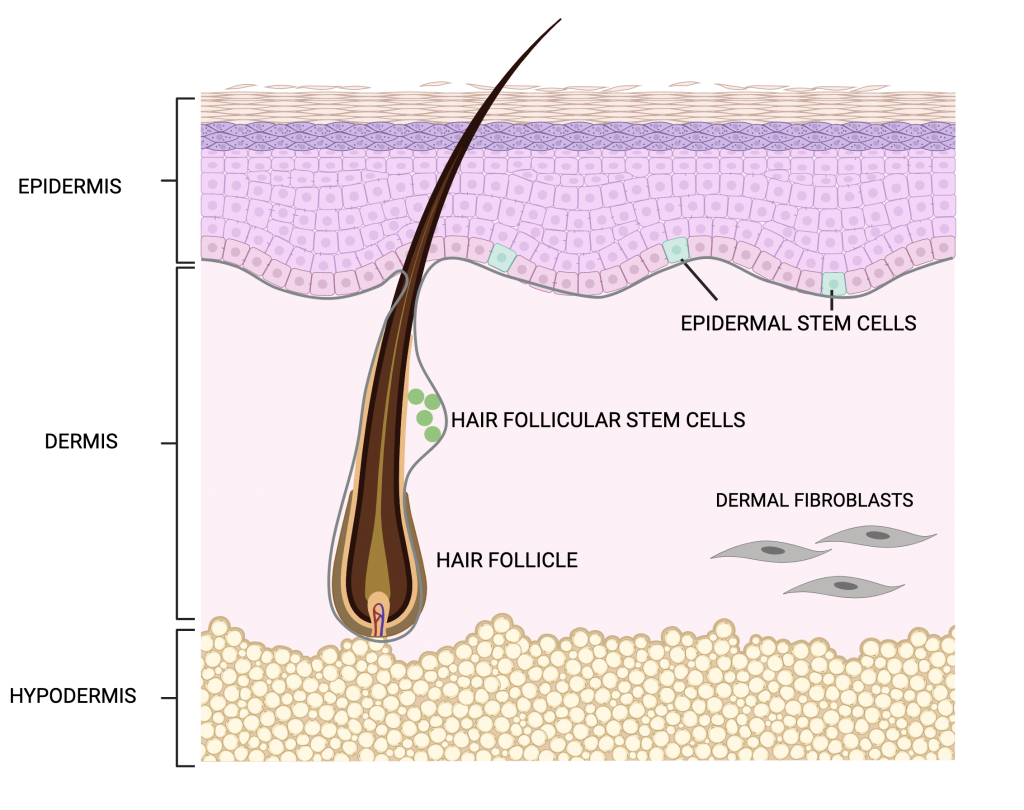
During inflammation, there is an influx of immune cells to clear invading microbes and cell debris at the site of injury. This is followed by the proliferation phase during which there is an increase in the production of epithelial cells that will migrate to the outer edge of the site of injury to repair the wound and restore it back to its uninjured state. An essential step in wound healing is the mobilization of stem cells for the formation of new epithelial cells. Skin stem cells are found in the basal layer of the epidermis and in the bulge area of hair follicles. Epidermal stem cells are actively involved in replenishing cells as skin undergoes normal homeostasis as well as during wound repair. Stem cells in the hair follicles, on the other hand have periodic patterns of rest and activity during hair growth. Following injury, however, hair follicular stem cells are also involved in rebuilding the epidermis to seal the open wound.
Interactions between the immune cells during the inflammation stage and stem cells during the proliferation stage of wound healing are important for efficient tissue repair to take place. Cytokines are signaling molecules produced by cells that are required for cell-cell communication to stimulate cell migration towards the site of injury. Molecular and cellular mechanisms to address the role of cytokines in mediating interactions between immune cells and stem cells during wound healing remain unexplained.
In a previous study, Pedro Lee and colleagues observed that mice that lacked the interleukin -1 receptor (IL-1R) for IL-1 cytokine had a delayed wound healing response. In the skin, IL-1 is released by damaged keratinocytes (keratin producing cells present in nails, hair and skin). and dysregulation of IL-1 has been associated with a number of skin diseases. Pedro Lee, Rupali Gund and colleagues conducted the current study to understand the mechanism behind delayed wound healing in IL-1R mutant mice. They analyzed the molecular and cellular interactions during wound healing using a genetically engineered mouse model in which the entire skin mimics the biological response of wound healing. This mouse model, which lacks the caspase 8 gene in the epidermis, exhibits a wound healing response even in the absence of injury, thereby providing the researchers with a large number of stem cells participating in a wound healing process. The authors studied the structure of tissues and gene expression patterns in the skin of these mice. The researcher additionally performed assays to analyze proliferation of cells by growing cells in the lab from the genetically engineered model and the mice lacking IL-1R. The researchers found that IL-1 mediates wound healing through activation of stem cell proliferation in two possible ways. The first is by activating dermal fibroblasts that will activate the epidermal stem cell to cover up the open wound (Fig.2.A). The second is the activation of a population of immune cells called gamma delta T(γδT)-cells. These cells in turn activate the resting stem cells found in the hair follicles. These activated stem cells then migrate from the hair follicle towards the site of the injury for wound healing. The researchers also found that IL-1 interacts with another cytokine, IL-7, and together they work to increase the number of active gamma delta T cells in wounded skin and secrete growth factors (Fig.2.B) thereby increasing the population of stem cells promoting wound healing.
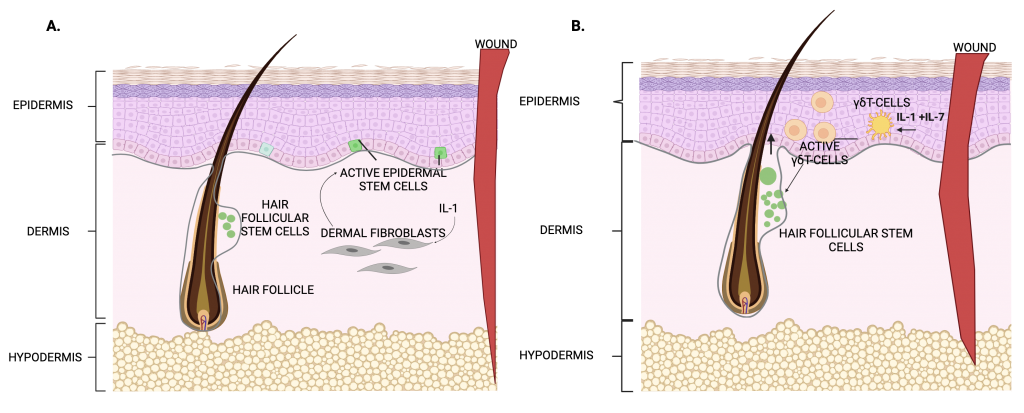
Normal wound healing is an important and complex physiological process to ensure timely healing to maintain skin integrity. It requires coordinated interactions between various factors, cells and cytokines at each healing stage. Lee, Gund and colleagues have identified a novel role for immune cells (γδT-cells) in tissue repair in addition to their well-established role of fighting infections. The ability of γδT cells to respond to IL-1 and, in turn, secrete growth factors that promote stem cell reparative activity expands the kind of functions that immune cells can perform in tissues in addition to their common role in immunity. Stem cell therapy and regenerative medicine shows great potential in the field of wound healing and skin regeneration.
This study reveals how immune cells communicate with stem cells during tissue repair and identifies new cellular interactions that can be targeted to prevent diseases in which wound healing is impaired. Such therapies will preclude need for invasive surgical interventions and skin graft procedures as a treatment for chronic wounds.
Dr. Rupali Gund is a postdoctoral research scientist in the department of dermatology at Columbia University Irving Medical Centre. Her research focuses on studying mechanisms in skin autoimmune diseases and finding new ways to design therapies to improve patient’s quality of life.