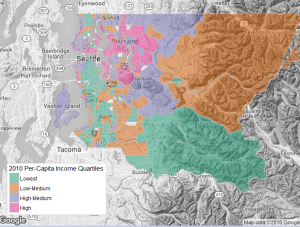
Per Capita Income in King County, WA.
Social Epi Cluster members Stephen Mooney and Gina Lovasi recently led an investigation into neighborhood and genetic contributions to cardiac arrest risk, finding that about 75% more cardiac arrest cases who had a high-risk genetic variant lived in socioeconomically deprived neighborhoods than would be expected due to chance alone. This finding, recently published ahead-of-print in Epidemiology highlights a potentially underappreciated way that precision medicine can inform population health: by helping to discriminate between theories regarding the causes of disease rather than solely by identifying more vulnerable subgroups of a population.
Sudden cardiac arrest – when someone’s heart stops beating regularly — is usually fatal, for obvious reasons. Cardiac arrest causes nearly 300,000 deaths per year in the United States, more commonly among people living in lower income neighborhoods. There are several reasons this might be. For one, people who live in poorer neighborhoods are usually themselves poorer, and poverty makes it harder to have a healthy diet or be sufficiently physically active, both of which protect against cardiac arrest. But individual poverty isn’t the only thing: disadvantaged neighborhoods also often have more polluted air, and exposure to air pollution is another known risk factor for cardiac arrest. And neighborhood conditions are also typically more stressful in lower income neighborhoods, and stress has been implicated as a risk factor for cardiac arrest as well. So where should we focus our cardiac arrest prevention efforts? It’s important to know which pathways are most implicated in cardiac arrest cases to know where to focus.
One way to identify the causes that can help focus population-based efforts is to study the most affected individuals. It turns out that cardiac arrest is somewhat more common among people with a common polymorphism in a gene called ADRB2, which produces proteins that help mediate responses to stress. Mr. Mooney and Dr. Lovasi hypothesized that if the stress pathway is a particularly important cause of cardiac arrest, then people with the higher-risk ADRB2 genetic variant should have even higher risk when living in lower income, disadvantaged neighborhoods (assuming disadvantaged neighborhoods had more stressful conditions owing to higher violence risks, less green space, etc.).
To explore this hypothesis, they examined neighborhood conditions and genetic variants using a case-only gene-environment interaction design with a dataset of adult cardiac arrest patients from King County, Washington, a large county whose core city is Seattle. They found that about 75% more cardiac arrest cases with the high-risk ADRB2 variant lived in the socioeconomically deprived neighborhoods than would be expected due to chance alone. Though preliminary, these findings nonetheless suggest that the stress pathway was an important cause of cardiac arrest in this population, and that there may be between-individual differences in vulnerability to stressful conditions.
To date, most discussions of the relationship between precision medicine and population health support using genetic data to identify particularly susceptible individuals. And indeed, that approach could be beneficial here, at least if there were appropriate interventions to apply those with the high risk ADRB2 variant. But the genetic data had another important use here in discriminating between theories regarding the causes of cardiac arrest. To understand this, consider that if air pollution were the primary cause of cardiac arrest in the subjects under study, then we would expect broadly higher risk among subjects living in disadvantaged neighborhoods, but not specifically disproportionately high risk among subjects with the high-risk ADRB2 variant. The fact that Mr. Mooney and Dr. Lovasi did see that disproportionately high risk supports the stress hypothesis. More generally, as we learn more about the human genome, using genetic data to distinguish between hypothesized causal mechanisms may help to improve population health.